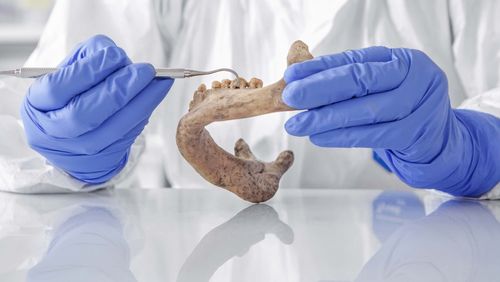
Antibiotics inventory in our mouths
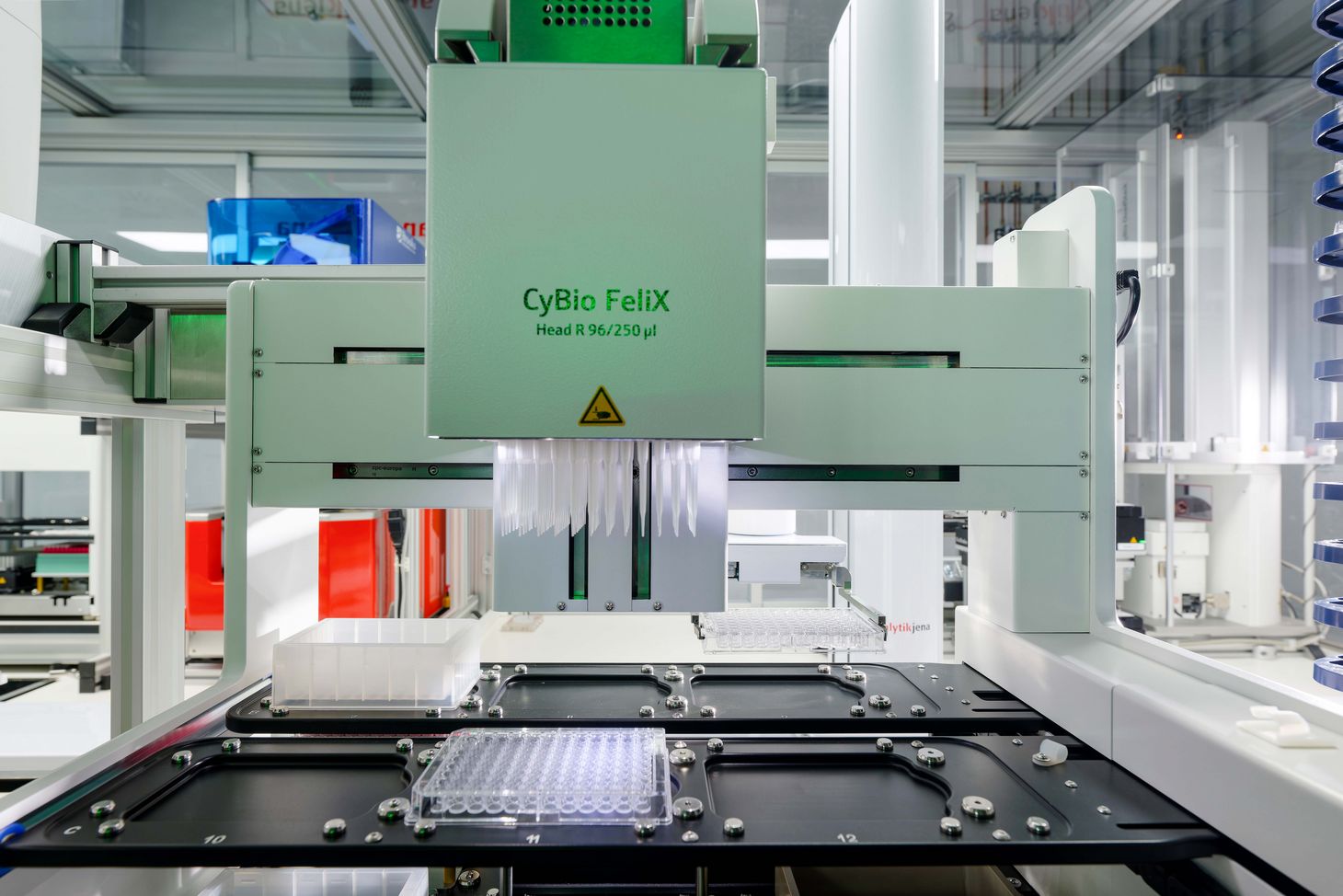
Dental calculus is a veritable “bioarchive” of human history. Recently, a team in the WSS palaeobiotechnology project has examined ancient and modern tooth tartar to document the evolution of antimicrobials and, for the first time ever, recreated one such potential antibiotic and tested its efficacy. A custom-made analysis platform made the work possible.
The human mouth is home to a complex microbial community in which hundreds of bacterial species compete with one another and, in doing so, maintain a symbiotic balance. A key role in this oral homeostasis is played by antimicrobial peptides (AMPs), small protein molecules used by bacteria to fight other microorganisms. In other words: AMPs are natural antibiotics. They’re also precisely the kind of molecules that are the focus of the palaeobiotechnology project in Jena, which the Werner Siemens Foundation (WSS) has financed for the past six years.
The research team led by chemist Pierre Stallforth and biomolecular archaeologist Christina Warinner is aiming to recreate antibiotic substances found from ancient skeletal remains—with the ultimate objective of developing drugs effective against modern resistant pathogens. The researchers are capitalising on the fact that the DNA of bacteria is particularly well preserved in dental calculus, also known as tooth tartar. In a widely acclaimed study published in Science three years ago, they showed for the first time that bacterial natural products could be recreated from the dental calculus of early humans living more than one hundred thousand years ago.
Since then, the team has made excellent progress, as demonstrated in a paper recently published in the Journal of the American Chemical Society (JACS). In their study*, the researchers used a bioinformatics pipeline they developed to directly and automatically identify antimicrobial peptides encoded in the bacterial DNA found in the dental calculus of early humans. And for the first time, they reconstructed ancient molecules that exhibit antimicrobial properties.
Systematic search through mountains of data
For the study, the researchers analysed the DNA of more than one hundred dental calculus samples stemming from present-day humans, archaeological humans and Neanderthals as well as from chimpanzees, gorillas and howler monkeys; the oldest samples were more than one hundred thousand years old. Finding the genetic sequences of antimicrobial peptides in the abundance of genetic material is, however, no straightforward task, as millions of potential candidates are hidden in the genomes. “Current identification tools generally yield too many false positives and false negatives,” Stallforth says.
To solve this problem, the research team engineered a new analysis tool called AMPcombi. It combines six existing tools, compares the results they generate, and then automatically filters out the peptides most likely to have functional antimicrobial properties. The filtering process ensures that candidate sequences match key criteria known to be necessary for effective antimicrobial properties. In the case of ancient gene sequences, the tool also analyses whether certain predictable forms of DNA damage known to occur over time are present, ensuring that the antimicrobial peptide sequences truly originate from ancient microorganisms.
Christina Warinner emphasises the importance of eliminating such “false alarms” at an early stage: “False positives slow down the research and significantly increase lab costs.” The new AMPcombi tool accelerates the identification of antimicrobial peptides by allowing the researchers to rapidly filter and categorise millions of potential candidates. “It allowed us to focus much more quickly and efficiently on the most promising AMP candidates for testing in the lab,” Warinner says.

Ancient active compounds redux
Using their new tool, the researchers identified seventy-eight gene clusters containing a total of one thousand six hundred and sixty-nine potential antimicrobial peptides, the overwhelming majority of which originated from the bacterial genus Actinomyces. These microorganisms have been a core component of the oral microbiome for at least forty million years, and they play a pivotal role in the formation of dental calculus. The antimicrobial peptides they produce are called “actifensins”.
The researchers compared the structure and genetic variants of these peptides, distinguishing between the “paleo-actifensins” from ancient samples and “modern actifensins” from present-day specimens. A fascinating pattern emerged: some of these molecules have remained virtually unchanged for tens of thousands of years, while others have changed significantly—apparently adapting their mode of action over time to new conditions in the oral cavity as well as to new microbial competitors. “It’s the first time that anyone has directly documented the development of an antimicrobial agent over the course of human evolution,” Warinner says.
To test whether the evolutionary changes also have functional implications, the researchers synthesised several actifensins using biotechnological methods, then tested them in the lab. And in fact, the synthesised actifensins exhibited antimicrobial properties. “We discovered that modern actifensins have a different activity spectrum than their ancient counterparts,” Stallforth explains. This, too, is a novel research finding: for the first time, antimicrobial peptides found in the dental calculus of early humans were “reanimated” and their efficacy was demonstrated. “That said, we don’t yet know which specific bacteria were the main targets of these ancient active compounds,” Warinner says.
Evolution of the oral microbiome
The study could provide a pathway to combating one of the biggest medical problems of our time: antibiotic resistance. An increasing number of bacteria have become resistant to common drugs, yet effective new medications are rarely developed. Antimicrobial peptides could prove a viable solution. Pierre Stallforth says the substances have various advantages. For example, they’re easy to modify, which would simplify the development of new variants to circumvent resistance. In addition, many antimicrobial peptides are known to attack bacterial cell walls and cell membranes. “This might force the bacteria to undergo significant changes to develop resistance,” Stallforth explains.
The work the palaeobiotechnology team is undertaking is also a journey into the past: to use antimicrobial peptides most efficiently, a deeper understanding of their evolution is critical. Stallforth says, “When we more fully understand how these molecules affect the oral microbial community, we can derive general principles that may deliver unexpected clues about how best to use antimicrobial peptides for therapeutic purposes.”
Last but not least, the two researchers say the study illustrates the importance of interdisciplinary collaboration. “The success of our work is based on close communication between bioinformaticians and scientists in the lab,” Pierre Stallforth says. Christina Warinner adds: “These are the kinds of research results and scientific advances that only happen when interdisciplinary teams work together on a daily basis.”
> Study